 J Mol Biol 215:403–410īader GD, Betel D, Hogue CW (2003) BIND: the biomolecular interaction network database. Bioinformatics 21:3596–3603Īltschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. Key pointsĪllen JE, Salzberg SL (2005) JIGSAW: integration of multiple sources of evidence for gene prediction. 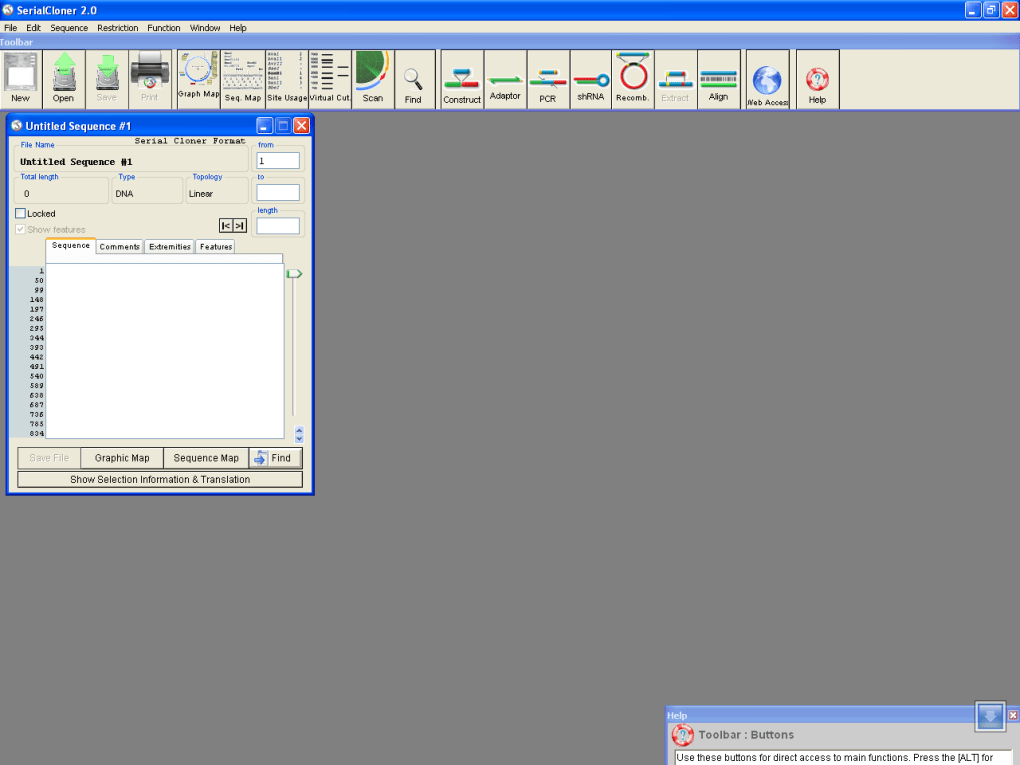 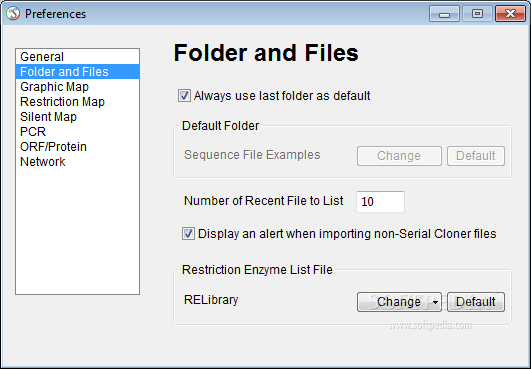
The intent of this article is to compile a list of tools and information resources used by scientists to treat information from the massive sequencing of recent platforms to new generations and the applications of this information in different areas of life sciences including medicine. Recently, sequencing platforms, a large scale of genomes and transcriptomes, have created new challenges not only to the genomics but especially for bioinformatics. The goal is to provide scientists with the right means to explain normal biological processes, dysfunctions of these processes which give rise to disease and approaches that allow the discovery of new medical cures. All this work produces an “ocean” of information that can only be “sailed” with the help of computerized methods. Bioinformatics combines the methods used in the collection, storage, identification, analysis, and correlation of this huge and complex information. With this menu entry, the user can map a single DNA sequence or collection of DNA fragments to any other DNA sample in the library or genome.While we have a basic understanding of the functioning of the gene when coding sequences of specific proteins, we feel the lack of information on the role that DNA has on specific diseases or functions of thousands of proteins that are produced. Another great feature is the menu entry "Serial Cloning Window". 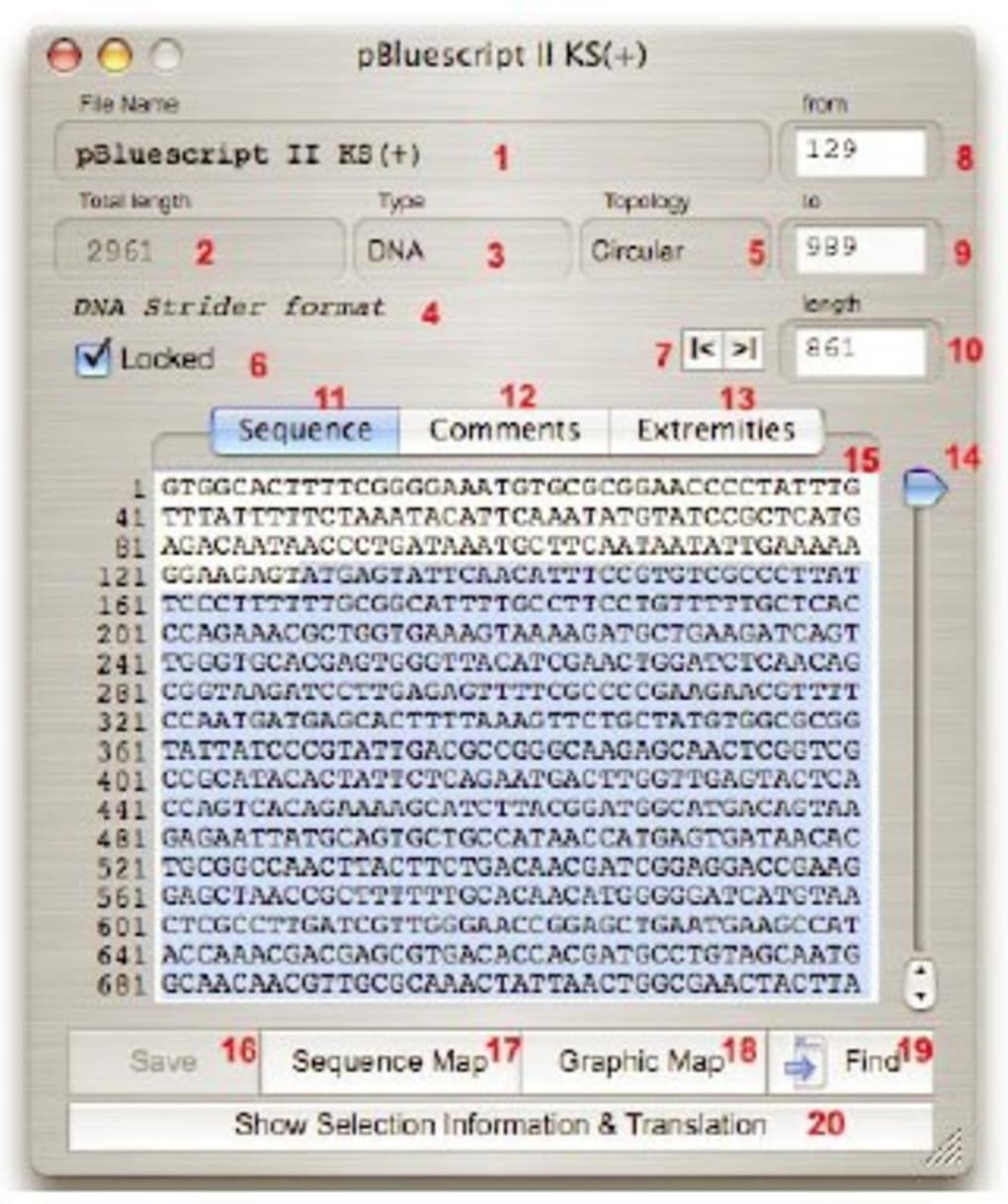
For example, there is a "Genetic Sequence Window" that contains an extensive list of genetic sequences that this software can automatically map and analyze. This is just a small sampling of the features that this great software program has to offer.Īnother great feature of this product is that users can manually select which windows and menus of the utility application they want to use to perform their tasks. The software has a large variety of useful functions such as real-time generation of mapped fragments, correction of errors in sequences, identification of individual genes, multipleogenicity detection, hybridization detection, population genetic analysis, scoring methods, and database support. The software is able to read and write DNA Strider-readable files and automatically import and exports data into the universal FASTA file format. The software has a graphical user interface (GUI) that makes sequencing and visualizing data easy and fast. The manual will show you the basic workflow that is involved with sequencing, cloning, and analyzing DNA samples. To give you an idea about how Serial Cloner works, let us take a look at the program's manual and how to use it. 
The software is available for purchase on the web and for download straight from the company's website. This software can be used by anyone with a Windows computer. Serial Cloner is an awesome software tool that allows users of molecular biology research to quickly and easily sequence, analyze, visualize, analyze, and interpret the data obtained during their experiments. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
